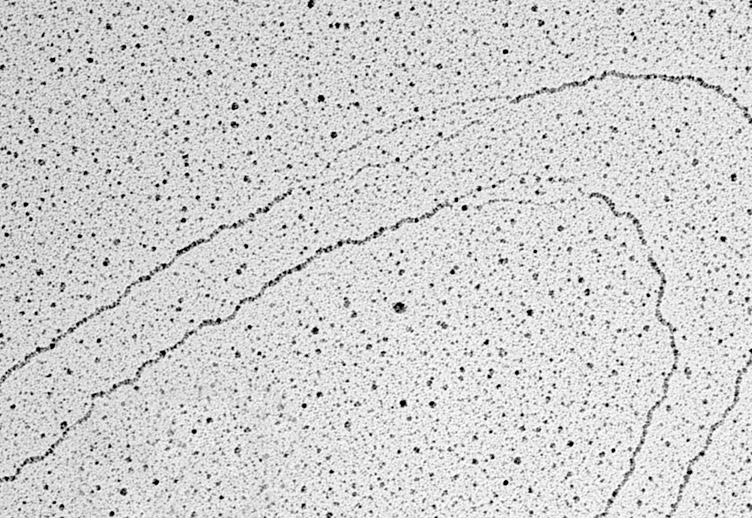
Replicating repeats: the checkpoint turns a blind eye
Commentary on Vincenzo Costanzo paper's published in Nature Cell Biology
October 2017
Half of the human genome is made up of repetitive DNA. However, mechanisms underlying replication of chromosome regions containing repetitive DNA are poorly understood. We reconstituted replication of defined human chromosome segments using bacterial artificial chromosomes in Xenopus laevis egg extract. Using this approach we characterized the chromatin assembly and replication dynamics of centromeric alpha-satellite DNA. Proteomic analysis of centromeric chromatin revealed replication-dependent enrichment of a network of DNA repair factors including the MSH2-6 complex, which was required for efficient centromeric DNA replication. However, contrary to expectations, the ATR-dependent checkpoint monitoring DNA replication fork arrest could not be activated on highly repetitive DNA due to the inability of the single-stranded DNA binding protein RPA to accumulate on chromatin. Electron microscopy of centromeric DNA and supercoil mapping revealed the presence of topoisomerase I-dependent DNA loops embedded in a protein matrix enriched for SMC2-4 proteins. This arrangement suppressed ATR signalling by preventing RPA hyper-loading, facilitating replication of centromeric DNA. These findings have important implications for our understanding of repetitive DNA metabolism and centromere organization under normal and stressful conditions.
[PMID 27111843]
We humans have a difficult time reading through texts with complicated character sequences and frequently stutter. A similar problem is faced by DNA Polymerases when they try to make a copy of DNA. Anyone with a bit of experience in Molecular Biology is well aware of this problem as amplifying repeated sequences by PCR is often problematic and leads to either failure of the reaction or to the presence of errors in the amplified copy. These problems observed in vitro, are nothing else but a recapitulation of the same problem that cells experience every S phase when trying to duplicate repetitive DNA.
While the key to cellular functions lies in genes, only 5-10% of our genome is made of genes or functional elements. The rest, frequently called as “dark-matter”, is to a large extent filled with repetitive sequences including leftovers of viral integrations, transposable elements and alike. Recent estimates indicate that up to two-thirds of the human genome may be present in the human genome. Interestingly, even if repeated sequences comprise most of our genome, they are often neglected in biomedical research studies, in part due to the technical difficulties in working with them. A paradigmatic example are most Next Generation Sequencing datasets, where the analysis starts by excluding repeated sequences since they cannot be mapped to a defined position in the human genome.
While the majority of the repetitive sequences might be non-functional and simply represent scars of past integrations of exogenous DNA, some of them play central roles in cellular biology such as telomeres, centromeres or ribosomal DNA. Consistent with the difficulties that DNA polymerases face during the replication of repetitive sequences, many of them are actually considered “fragile sites”. The instability of ribosomal DNA is well known particularly from yeast studies, and has been even associated to the process of ageing. Telomeres have also recently been found to be fragile, and they contain a unique dedicated pathway to complete their replication. To what extent the replication of other repeats, including centromeres or rDNA, also demands ad-hoc machinery, remains unknown.
To tackle this problem, the group of Vincenzo Costanzo took an original approach that exploited the usefulness of frog oocyte extracts to replicate exogenous DNA. Antoine Aze and colleagues explored how Bacterial Artificial Chromosomes (BACs) containing repeats from human centromeric alpha-satellite sequences were replicated in these extracts, and compared it to the replication of BACs containing a similar GC base content but free of repeats. Their strategy proved to very useful as it yielded important insights into the replication or centromeric DNA.
The first cool observation was that human BACs added to Xenopus extracts form a nucleus, similar to the endogenous one. The BAC containing centromeric repeats (cenBAC) was then replicated, albeit more slowly than the control BAC (cBAC). Subsequent proteomic analysis identified proteins that were enriched (or depleted) from the replicating BACs. These analyses revealed that cenBACs presented a higher presence of DNA repair factors, which might be there as a consequence of DNA breaks arising during DNA replication, or perhaps are already there in a preemptive position “just in case” breaks do happen. Particularly enriched were components of the Mismatch Repair machinery, which would make sense since these factors travel with the replisome to correct mistakes placed by DNA polymerases. It is also possible, however, that the increase in MMR proteins simply reflects a higher amount of replisomes in the cenBACs, which could be due to the more frequent stalling of the replication forks. Other factors enriched in replicating cenBACs were proteins involved in chromosome architecture and topology such as Topoisomerases or components of SMC complexes, likely due to the particular topology of centromeric chromatin.
A surprise, however, was to find that several components from the replication checkpoint were depleted from cenBACs. In vertebrates, the S phase checkpoint is coordinated by the ATR kinase. The activation of ATR is mediated by the ssDNA-binding factor RPA, which together with the recruitment of additional factors such as the allosteric activator TOPBP1, trigger the kinase activity of ATR. In other words, what the checkpoint first “smells” is actually the accumulation of ssDNA at stalled replication forks. In this context, and as readily imagined by the Costanzo team, one mechanism to explain why checkpoint factors could be depleted from centromeric DNA is that the abundance of ssDNA is somewhat limited at these sequences. And this is exactly what they found.
Using another technique mastered by their team, Electron Microscopy, Aze and colleagues found that replicating cenBACs were full of looped sequences. The most parsimonious interpretation is that when the double helix opens up during DNA replication, the repetitive nature of the sequences will lead to their spontaneous formation of loops and other sort of secondary structures, thus occluding the presence of ssDNA and the activation of the checkpoint. In support of this, the use of Topoisomerase I inhibitors reduced the presence of supercoils and restored checkpoint activation in centromeric DNA.
The story is round and provides one of the first examples and initial insights as to how centromeric DNA is replicated. The discovery that the checkpoint is somewhat silenced at this region is surprising, but makes sense from an evolutionary point of view. ATR activates the alarm when DNA replication problems occur, leading to cellular consequences that can include the activation of apoptosis. Since repetitive sequences will always face problems during their replication, one could envision that the bar to activate the checkpoint should be a bit higher at these regions. In other words, the checkpoint needs to turn a blind eye at repetitive regions.
Finally, as any important study, this work also brings to mind many new questions that could be now tackled. For instance, what about the new and uncharacterized factors that were found in the proteomic studies as enriched in replicating cenBACs? Is it possible that, like in telomeres, the replication of centromeres also uses specialized machinery? Additionally, the group could also exploit this system to address additional questions about centromere replication. I am a good friend of Vincenzo, who I consider one of the most original scientists from our field. I still remember one of our conversations where he told me that he believes that Homologous Recombination factors are not essential in yeast, but yes in mammals, due to the higher presence of repeats in the mammalian genome. I have always liked the idea! They now have the opportunity to address this experimentally. To start with, they could simply explore how the depletion of HR proteins affects the replication of repetitive DNA. Anyway, from what I know of the team, I am sure that I cannot really predict what will be next from their lab. What I do know is that it will help all of us to understand a bit more how life works.